Note
Go to the end to download the full example code or to run this example in your browser via Binder.
Interact with 3D images (of kidney tissue)#
In this tutorial, we explore interactively a biomedical image which has three
spatial dimensions and three color dimensions (channels).
For a general introduction to 3D image processing, please refer to
Explore 3D images (of cells).
The data we use here correspond to kidney tissue which was
imaged with confocal fluorescence microscopy (more details at [1] under
kidney-tissue-fluorescence.tif).
import matplotlib.pyplot as plt
import numpy as np
import plotly
import plotly.express as px
import skimage as ski
Load image#
This biomedical image is available through scikit-image’s data registry.
data = ski.data.kidney()
The returned dataset is a 3D multichannel image:
print(f'number of dimensions: {data.ndim}')
print(f'shape: {data.shape}')
print(f'dtype: {data.dtype}')
number of dimensions: 4
shape: (16, 512, 512, 3)
dtype: uint16
Dimensions are provided in the following order: (z, y, x, c), i.e.,
[plane, row, column, channel].
n_plane, n_row, n_col, n_chan = data.shape
Let us consider only a slice (2D plane) of the data for now. More
specifically, let us consider the slice located halfway in the stack.
The imshow function can display both grayscale and RGB(A) 2D images.
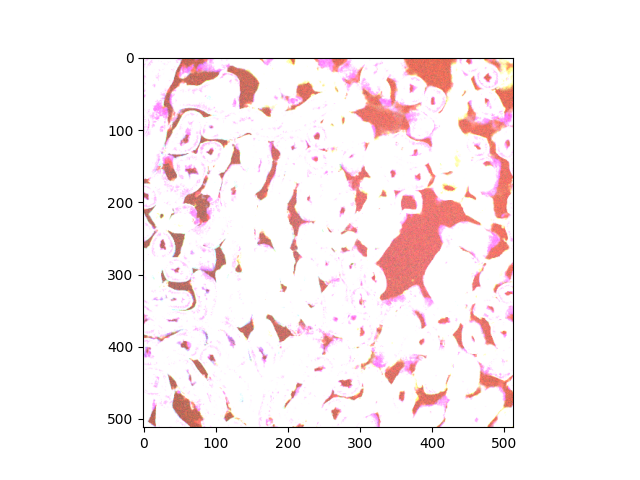
Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers). Got range [64..4095].
<matplotlib.image.AxesImage object at 0x7ff901d3bce0>
According to the warning message, the range of values is unexpected. The image rendering is clearly not satisfactory color-wise.
range: (10, 4095)
We turn to Plotly’s implementation of the plotly.express.imshow() function, for it
supports value ranges
beyond (0.0, 1.0) for floats and (0, 255) for integers.
Here you go, fluorescence microscopy!
Normalize range for each channel#
Generally speaking, we may want to normalize the value range on a per-channel basis. Let us facet our data (slice) along the channel axis. This way, we get three single-channel images, where the max value of each image is used:
What is the range of values for each color channel? We check by taking the min and max across all non-channel axes.
range for channel 0: (10, 4095)
range for channel 1: (68, 4095)
range for channel 2: (35, 4095)
Let us be very specific and pass value ranges on a per-channel basis:
Plotly lets you interact with this visualization by panning, zooming in and out, and exporting the desired figure as a static image in PNG format.
Explore slices as animation frames#
Click the play button to move along the z axis, through the stack of all
16 slices.
